Sheng Li, Ph.D.
Associate Professor
Applies data integration and machine learning to advance the frontiers of cancer epigenomics and evolution.
Visit The Li Lab
Location
My research interest is to understand the inner workings of cancer cells – the genetic and epigenetic heterogeneity that drive cancer initiation and progression. We utilize computational and sequencing methodologies to identify and characterize the essential epigenetic lesions that guide cancer cells to evolve and escape from anti-cancer therapy. The ultimate goal is to develop novel methods to predict and address tumor evolution.
Selected Publications
- 72 Katebi A, Chen X, Ramirez D, Li S, Lu M. Data-driven modeling of core gene regulatory network underlying leukemogenesis in IDH mutant AML. NPJ Syst Biol Appl. 2024 Apr 9; 10(1):38
- 41 Zhao Y, Agyemang D, Liu Y, Mahoney M, Li S. Quantifying Interpretation Reproducibility in Vision Transformer Models with TAVAC. bioRxiv. 2024 Jan 22
- 15 Anczuków O, Airhart S, Chuang JH, Coussens LM, Kuchel GA, Korstanje R, Li S, Lucido AL, McAllister SS, Politi K, Polyak K, Ratliff T, Ren G, Trowbridge JJ, Ucar D, Palucka K. Challenges and opportunities for modeling aging and cancer. Cancer Cell. 2023 Apr 10; 41(4):641-645
- 0 Li S. Inferring the Cancer Cellular Epigenome Heterogeneity via DNA Methylation Patterns. Cancer Treat Res. 2023; 190:375-393
- 71 Antel M, Raj R, Masoud MYG, Pan Z, Li S, Mellone BG, Inaba M. Interchromosomal interaction of homologous Stat92E alleles regulates transcriptional switch during stem-cell differentiation. Nat Commun. 2022 Jul 9; 13(1):3981
- 106 Liu Y, Rosikiewicz W, Pan Z, Jillette N, Wang P, Taghbalout A, Foox J, Mason C, Carroll M, Cheng A, Li S. DNA methylation-calling tools for Oxford Nanopore sequencing: a survey and human epigenome-wide evaluation. Genome Biol. 2021 Oct 18; 22(1):295
- 23 Chen X, Ashoor H, Musich R, Wang J, Zhang M, Zhang C, Lu M, Li S. epihet for intra-tumoral epigenetic heterogeneity analysis and visualization. Sci Rep. 2021 Jan 11; 11(1):376
- 83 Malkani S, Chin CR, Cekanaviciute E, Mortreux M, Okinula H, Tarbier M, Schreurs AS, Shirazi-Fard Y, Tahimic CGT, Rodriguez DN, Sexton BS, Butler D, Verma A, Bezdan D, Durmaz C, MacKay M, Melnick A, Meydan C, Li S, Garrett-Bakelman F, Fromm B, Afshinnekoo E, Langhorst BW, Dimalanta ET, Cheng-Campbell M, Blaber E, Schisler JC, Vanderburg C, Friedländer MR, McDonald JT, Costes SV, Rutkove S, Grabham P, Mason CE, Beheshti A. Circulating miRNA Spaceflight Signature Reveals Targets for Countermeasure Development. Cell Rep. 2020 Dec 8; 33(10):108448
- 50 Li S, Chen X, Wang J, Meydan C, Glass JL, Shih AH, Delwel R, Levine RL, Mason CE, Melnick AM. Somatic Mutations Drive Specific, but Reversible, Epigenetic Heterogeneity States in AML. Cancer Discov. 2020 Dec; 10(12):1934-1949
- 51 Li P, Lu M, Shi J, Gong Z, Hua L, Li Q, Lim B, Zhang XH, Chen X, Li S, Shultz LD, Ren G. Lung mesenchymal cells elicit lipid storage in neutrophils that fuel breast cancer lung metastasis. Nat Immunol. 2020 Nov; 21(11):1444-1455
- 90 Zhao Y, Pan Z, Namburi S, Pattison A, Posner A, Balachander S, Paisie CA, Reddi HV, Rueter J, Gill AJ, Fox S, Raghav KPS, Flynn WF, Tothill RW, Li S, Karuturi RKM, George J. CUP-AI-Dx: A tool for inferring cancer tissue of origin and molecular subtype using RNA gene-expression data and artificial intelligence. EBioMedicine. 2020 Nov; 61:103030
- 58 Tong L, Wu PY, Phan JH, Hassazadeh HR, SEQC Consortium, Tong W, Wang MD. Impact of RNA-seq data analysis algorithms on gene expression estimation and downstream prediction. Sci Rep. 2020 Oct 21; 10(1):17925
-
 Featured ArticleOctober 24, 2023
Featured ArticleOctober 24, 2023Applying AI and machine learning to complex research tasks
JAX researchers are at the forefront of developing and implementing AI-based technologies for a variety of research applications. -
 Profile StoryNovember 09, 2021
Profile StoryNovember 09, 2021Using next-gen technology to explore how aging raises the risk for leukemia
JAX's Sheng Li is exploring how aging raises the risk for leukemia, and works to improve cutting edge tools for data analysis. -
 Research HighlightJanuary 12, 2021
Research HighlightJanuary 12, 2021Few mutations, important difference in cancer cells
JAX Professor Sheng Li, Ph.D., presents a tool, epihet, that automates the epigenetic analysis of AML cells. epihet allows researchers to assess epigenetic differences between cells, identify genomic locations where the differences occur, and associate cellular epigenetic profiles with cancer-related biological function. -
 Research HighlightJune 17, 2020
Research HighlightJune 17, 2020New research reveals the role of epigenetic dysfunction in blood cancer
A new discovery by JAX cancer researcher Sheng Li points to the possibility of new lymphoma treatments that would restore the protective effect of TET2 in tumors where it has been mutated.
-
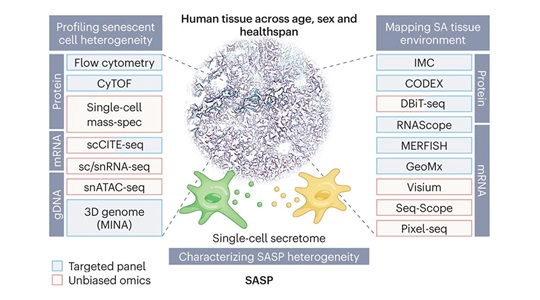 Research HighlightDecember 20, 2022
Research HighlightDecember 20, 2022Revealing the biology of senescent cells
JAX is playing a major role to bridge experimental mouse and human data and reveal the biology of senescent cells. -
 Research HighlightJanuary 18, 2021
Research HighlightJanuary 18, 2021JAX uses genetic models to fight CUP
A JAX team developed CUP-AI-Dx, a machine learning tool that uses RNA sequence data for analysis. The researchers show that CUP-AI-Dx provides an important clinical tool to help guide therapies for CUP patients. -
 Research HighlightOctober 20, 2020
Research HighlightOctober 20, 2020Connecting genetics with epigenetics in acute myeloid leukemia
Acute myeloid leukemia (AML) is an aggressive, difficult-to-treat cancer. Nonetheless, researchers have found that there are not as many genetic mutations associated with AML as with most other cancers. Aberrant epigenetic patterning, on the other hand, is emerging as a hallmark of AML. -
 Press ReleaseMarch 10, 2020
Press ReleaseMarch 10, 2020Sheng Li named an AACR NextGen Star
Jackson Laboratory (JAX) Assistant Professor Sheng Li, Ph.D., was chosen as a 2020 NextGen Star by the American Association for Cancer Research (AACR). She is one of 12 early-career cancer researchers recognized for their outstanding work and future potential.