Investigates how alternative RNA splicing contributes to cancer with the goal of identifying novel clinical biomarkers and targets for precision medicine.

Investigates how alternative RNA splicing contributes to cancer with the goal of identifying novel clinical biomarkers and targets for precision medicine.
The long-term goals of the Anczuków Lab are to define how rewiring of the splicing machinery contributes to tumor initiation, progression, and drug response, what triggers splicing alterations in tumors, and ultimately to translate this knowledge to develop innovative RNA-targeted therapeutics.
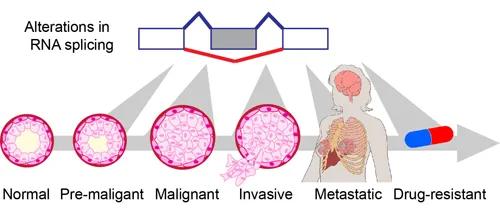
Figure1. Misregulation of alternative splicing contributes to various steps of cell transformation.
Building on our strengths in RNA and cancer biology, our lab research currently focuses on: 1) Targeting the regulation of splicing factors in cancer; 2) Defining aging-associated splicing as an oncogenic factor in breast; 3) Developing approaches to functionally characterize spliced isoforms at scale.
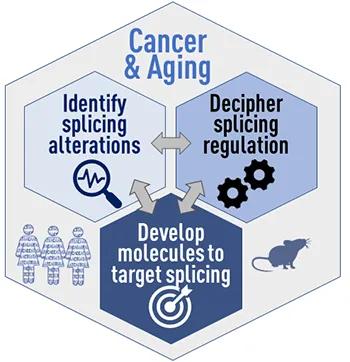
Figure 2. Our research program will uncover new RNA-driven mechanisms, biomarkers, and drug targets for human cancers, including those associated with aging.