Julia Oh, Ph.D.
Associate Professor
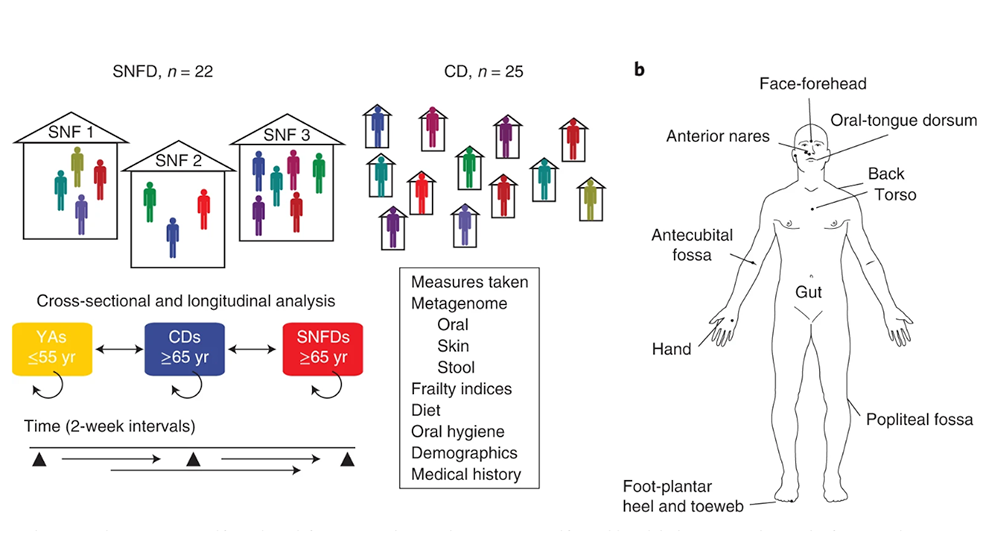
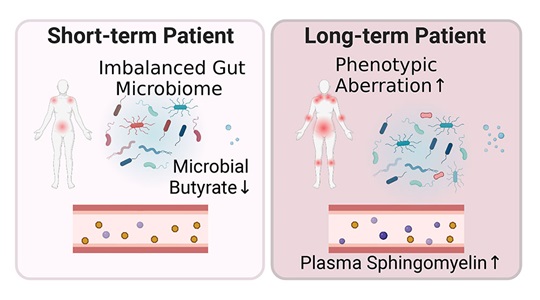
Our central goal is to develop microbiome therapeutics to treat human disease. We use diverse tools like genomics and synthetic biology to investigate our microbiome’s role in our health and engineer therapeutics.
Our central goal is to develop microbiome therapeutics to treat human disease. We use diverse tools like genomics and synthetic biology to investigate our microbiome’s role in our health and engineer therapeutics.
Education and Experience
Associate Professor, The Jackson Laboratory for Genomic Medicine
Postdoc, National Human Genome Research Institute, National Institutes of Health
Ph.D., Stanford University
B.A., Harvard University