When the superbug uprising occurs, will we be prepared?
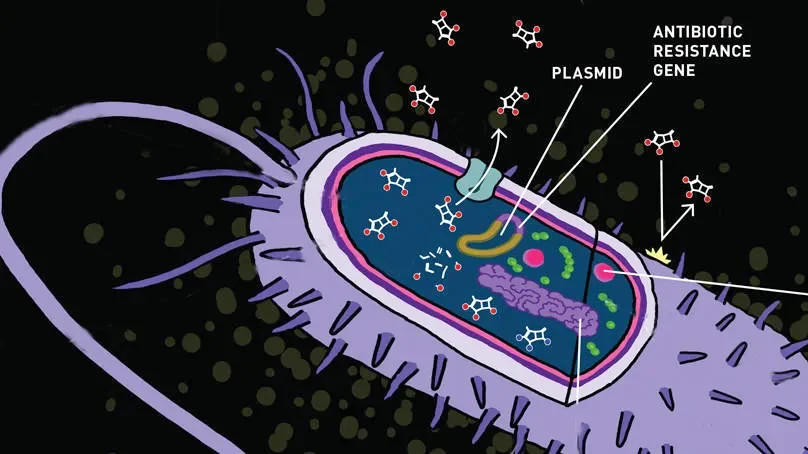
Superbug Invasion: coming soon, to a neighborhood near you
In May 2016, and for the first time in the United States, a colistin-resistant strain of E. coli, or “superbug,” was cultured from a patient’s urine sample. Superbugs are bacteria resistant to most prescribed antibiotics including “last resort” antibiotics such as colistin. Superbugs are thought to arise in response to inappropriate, widespread antibiotic use in agriculture and in the clinic. For example, not taking an antibiotic as directed, may result in incomplete elimination of a bacterial infection, leading to survival of strains of bacteria that evolve to resist that antibiotic.
Only months before the U.S. finding, routine E. coli cultures from food animals in China uncovered the presence of a colistin resistance gene, which could move from one strain of bacteria to another in a process called horizontal gene transfer within DNA plasmids. This gene has now spread to multiple species of bacteria and it has been detected in several countries. Prior to this study, the only known means to develop colistin resistance was through molecular changes within the chromosome of a single bacterium, and those mutations could only be transmitted to its descendants. As the American patient had not traveled abroad, finding colistin-resistant bacteria in the U.S. was, and remains, a concerning discovery.
Undoubtedly this is not the first time you have read about antibiotic resistant bacteria. Reports like these tend to gather media attention for a short period of time, and are then quietly ignored until the next big superbug infection discovery. For those in the infection and immunity research community, however, antibiotic resistance is a major global health problem.
Antibiotic resistance worldwide
Antibiotic resistant bacteria require significant medical and financial resources to treat and contain. For example, the U.S. Centers for Disease Control and Prevention (CDC) estimates that the annual impact of antibiotic-resistant infections in the US alone is $20 billion in direct health care costs. Methicillin-resistant Staphylococcus aureus (MRSA), untreatable gonorrhea, and extensively drug-resistant tuberculosis (XDR TB) top the list of the most significant threats worldwide.
Organizations such as the CDC, the World Health Organization (WHO), and the European Centre for Disease Prevention and Control (ECDC) recommend and implement measures to limit antibiotic resistance. Common themes across these agencies include improving health monitoring and disease detection, increasing sanitation practices while reducing overall antibiotic use, educating the public on appropriate antibiotic use, and encouraging research and development of novel therapeutics through funding incentives.
Mechanisms of antibiotic resistance
In general, misuse of antibiotics generates populations of resistant bacteria. Many of the drugs currently used clinically focus on targeting a very specific component in the bacterial life cycle, such as cell wall, nucleic acid, or protein synthesis pathways. Because bacteria have short replication times (on the order of 30-60 minutes per generation), bacteria have the ability to evolve rapidly. Additionally, bacteria can transfer genetic information with each other, as was demonstrated for the colistin resistance gene, through transformation, transduction, or conjugation. Taking these points together, if a single bacterium has a survival advantage against a particular antibiotic, such as by pumping the drug out of the cell, inactivating the drug, or using alternative methods to survive a drug targeted at a particular protein, a resistant infection can develop quickly and spread to other bacterial strains easily.
Source: CDC
Where have all the drugs gone?
Drugs for resistant bacteria are rare. In the 1950’s and 1960’s antibiotics were being developed regularly. Currently, out of 84 drugs approved by the US Food and Drug Administration (FDA) in 2015, two drugs were indicated for general antibiotic purposes. The small number of antibiotics in the drug pipeline by Pharma may be a result of the relatively low return on investment compared to drugs for cancer, cardiovascular disease, or diabetes. Executive actions by the Obama Administration that emphasize and incentivize developing therapies for antibiotic resistant infections may help correct this imbalance.
Antibiotic development strategies
In the U.S., the President’s Council of Advisors on Science and Technology (PCAST) is particularly interested in understanding mechanisms in antibiotic resistance and novel antibiotic discovery strategies. Some of these strategies are being examined in the lab.
For example, some therapies focus on re-sensitizing bacteria to antibiotics they are resistant to by co-treatments with novel inhibitors that complement or synergize their activity. The proposed mechanism in this study suggested that MRSA may become susceptible to β-lactam antibiotics, which prevent cell wall biosynthesis, by combining them with new drugs that compromise bacterial cell wall integrity. Working together, these drug combinations may overcome some forms of antibiotic resistance.
Tamoxifen, which is commonly used in treating breast cancer, may be repurposed to stimulate neutrophils to kill microbes like MRSA. This form of immunotherapy, in which the normal immune system becomes better at recognizing or acting against its natural targets, is predicted to improve efficacy against antibiotic resistant bacteria because the drug works indirectly, rather than directly, on bacterial populations capable of evading particular drugs. Another advantage to this approach is that tamoxifen is already approved for use in people, which would shorten the FDA approval process.
Other antibiotic treatments using ideas borrowed from cancer therapy research include uncovering MRSA hiding dormant within macrophages. This approach specifically directs antibiotics to target cells by conjugating them with infection-specific antibodies. These antibody-mediated therapies demonstrate a “narrow spectrum” approach, in that antibacterial activity is directed only at afflicted cells, rather than broadly killing systemic commensal bacteria, such as the naturally occurring microbiome.
With regards to the microbiome, some researchers are taking a basic biology approach to better understand how early and potent antibiotic use affects long-term bacterial profiles and an individual’s physiology. This group demonstrated that early antibiotic use reduces host gut microbiome diversity, potentially contributing to future adiposity.
Hopefully, infectious disease researchers may be able to take advantage of these global action plans from WHO, CDC, ECDC, or other health agencies to fund novel antibiotic development before superbug infections pervade strongly. In the meantime, U.S. Federal Agencies are taking the emergence of this resistance gene very seriously and are coordinating efforts to try to prevent its spread.
Suggested tools at The Jackson Laboratory to expedite infectious disease research:
JAX whitepaper on infectious disease modeling using humanized NSG mice
