Learn how three powerful CRISPR/Cas9 approaches work to edit mouse genomes and generate pre-clinical models for developing and validating therapies for human diseases.
Three CRISPR Approaches for Mouse Genome Editing
Blog Post | February 15, 2016
In the few short years since Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) and CRISPR-associated proteins (Cas) were first described, their use in editing the mouse genome has exploded. The CRISPR/Cas9 system is a tremendously versatile tool for engineering a wide variety of genetic alterations including knockouts (KO), deletions, point mutations, and short insertions (e.g. loxP sites, epitope tags, etc.). As an early adopter of the CRISPR/Cas9 technology, The Jackson Laboratory has been exploring the technique’s capabilities and limitations for generating mouse models.
The CRISPR-Cas9 system offers significant advantages over more traditional, directed mutagenesis methods, including lower costs, shorter timelines, and the capacity to alter multiple genes simultaneously. CRISPR/Cas9 gene editing is a rapidly growing field, and new techniques and methods are evolving accordingly. To date, three powerful CRISPR/Cas9 approaches to edit mouse genomes have been demonstrated:
1. Mouse embryos: The classic CRISPR approach
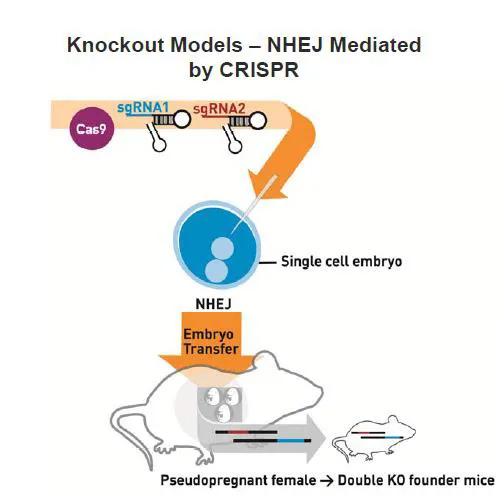
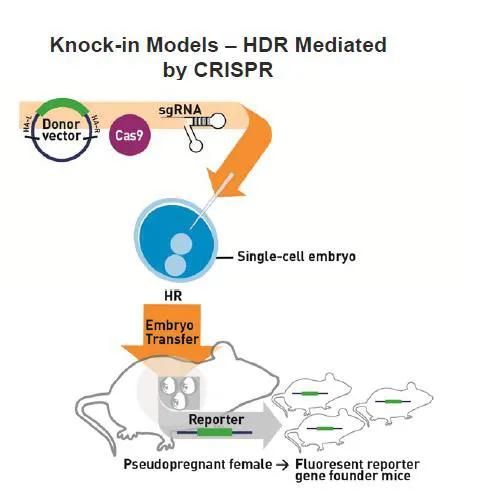
New mouse models can be generated with CRISPR/Cas9 by injecting Cas9 mRNA and either one or multiple single guide RNAs (sgRNA) directly into mouse embryos to generate precise genomic edits into specific loci (Figures 1 and 2). Mice that develop from these embryos are genotyped or sequenced to determine if they carry the desired mutation(s), and those that do are bred to confirm germline transmission. As CRISPR/Cas9 will work in most mouse strains, new mutations can be directly generated in a genetic background of choice. This eliminates the need, time, and resources to backcross mutations from one genetic background to another, which is a typical practice when generating mutant mice by traditional methods.
New mutations also can be added to existing mouse strains that already carry desired mutations, reducing the time and costs to generate double and triple mutant mice. To date, JAX has used CRISPR/Cas9 to successfully generate more than 110 KO models (both indel and deletion) and 69 knock-in models (point mutations, amino acid tags, LoxP sites and reporter insertions, as shown in Table 1).
2.Cas9-expressing mice
Generating mutations via direct Cas9 mRNA injection into embryos requires the equipment and expertise for embryo microinjection, which may not be readily available to all researchers.
Gene editing using CRISPR/Cas9, also can introduce mutations into specific organs or tissues in postnatal mice.
However, due to limitations in the size of constructs that can be efficiently inserted using current CRISPR/Cas technologies, complex projects including, sgRNA expression cassettes, promoters, polyadenylation, and fluorescent protein sequence cannot all be incorporated into a single viral expression vector that also includes Cas9 cDNA.
To overcome this obstacle, mouse strains with Cre recombinase-dependent Cas9 expression have been generated. These mouse strains allow forin vivo CRISPR gene editing wherever a viral vector co-expressing Cre and the sgRNA is injected. The virally-expressed Cre turns on Cas9 expression, which in turn edits the targeted gene or genes. Recent studies have successfully used this technique to edit the mouse genome in the brain, lungs, and pancreas with high efficiency. Cas9-expressing mice can dramatically accelerate our understanding of pathological processes that involve multiple genes or mutations, like tumor development.
Use of Cas9-expressing mice in this manner reduces the time and resources it would take to design traditional gene-targeted mutants and to cross these multiple mutants together to create mice with the necessary complex genotypes. Mice that express Cas9 systemically can be used for similar studies. With these mice, only injections with a sgRNA- expressing virus is required.
JAX distributes many Cas9 expressing mice, including the B6J.129(B6N)-Gt(ROSA)26Sortm1(CAG-cas9*,-EGFP)Fezh/J (026175) and B6J.129(Cg)-Gt(ROSA)26Sortm1.1(CAG-cas9*,-EGFP)Fezh/J (026179) mouse strains, which are on a C57BL/6J genetic background and express Cas9 in cre dependent and systemic fashions, respectively.
3. Cas9 and sgRNA expressing viruses
In vivo gene editing in mice also can be accomplished by local or systemic injection of separate Cas9 and sgRNA expressing lenti- or adeno-associated viruses.
CRISPR/Cas9-mediated gene editing occurs in cells that express both expression vectors. This approach has proved successful in disruptingmultiple genes simultaneously in neurons and in editing the Dmdmdx mutation in muscle, thereby restoring dystrophin protein expression and partially alleviating the dystrophic muscles’ functional deficiencies. Potentially any inbred, mutant, or transgenic mouse strain can be genetically modified using this technique, too, opening up nearly any established mouse model to precise genome editing.
With these variations to enlist CRISPR gene-editing technologies, we can quickly engineer mouse models to carry the same mutations present in patients with specific genetic diseases, and use them as pre-clinical models for developing and validating novel therapies.
Do you need a gene edited mouse model fast but don’t have the expertise yet? Get started by contacting ourTechnical Support Scientists to discuss your project.
