For in-depth product & services help, ask our
Technical Information Scientists
Stock No: 016198
Protocol 26962: Standard PCR Assay - Tg(tTA)
Version 5.2
Notes
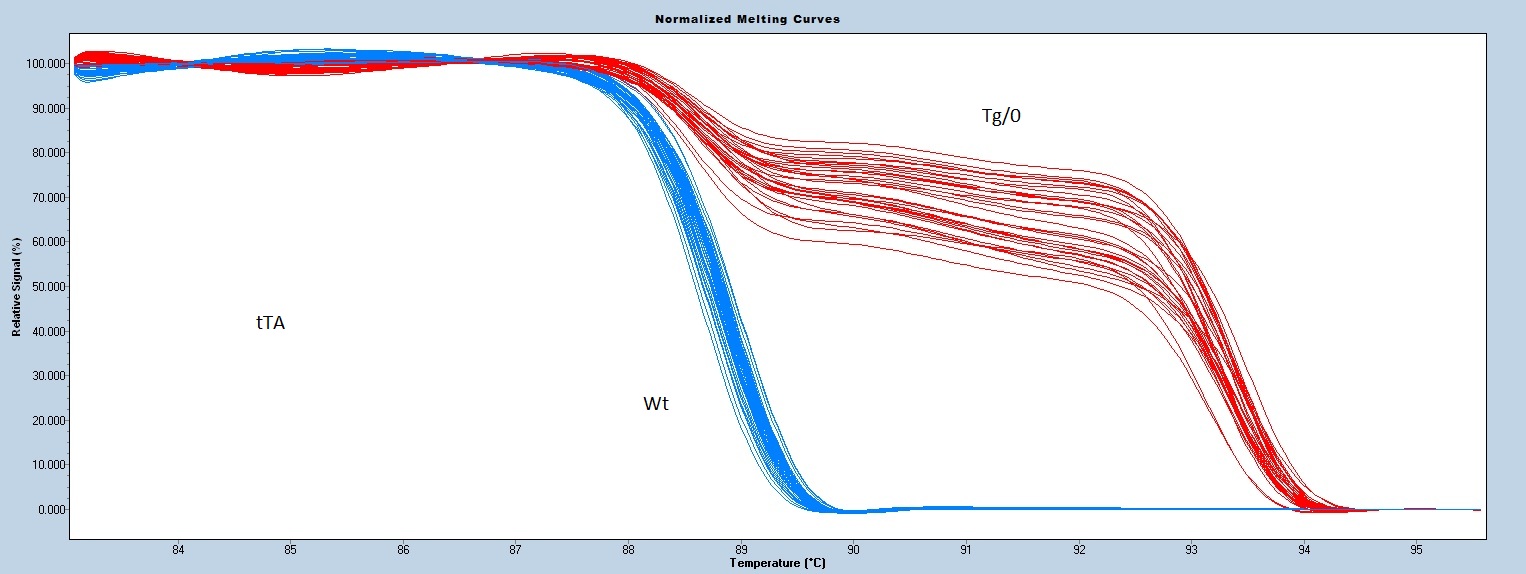
Melting curve analysis is done using a Roche Light Cycler 480.
This assay will NOT distinguish hemizygous from homozygous transgenic animals.
The genotyping protocol(s) presented here have been optimized for reagents and conditions used by The Jackson Laboratory (JAX).
To genotype animals, JAX recommends researchers validate the assay independently upon receipt of animals into their facility.
Reaction cycling temperature and times may require additional optimization based on the specific genotyping reagents used.
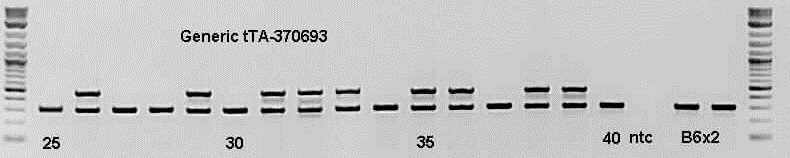
Expected Results
Transgene = ~450 bp
Internal positive control = 297 bp
JAX Protocol
Protocol Primers
| Primer | 5' Label | Sequence 5' → 3' | 3' Label | Primer Type | Reaction | Note |
|---|---|---|---|---|---|---|
| oIMR8746 | CGC TGT GGG GCA TTT TAC TTT AG | Transgene Forward | A | |||
| oIMR8747 | CAT GTC CAG ATC GAA ATC GTC | Transgene Reverse | A | |||
| oIMR9020 | AAG GGA GCT GCA GTG GAG TA | Wild type Forward | A | |||
| oIMR9021 | CCG AAA ATC TGT GGG AAG TC | Wild type Reverse | A |
Reaction A
| Component | Final Concentration |
|---|---|
| ddH2O | |
| Kapa 2G HS buffer | 1.30 X |
| MgCl2 | 2.60 mM |
| dNTP KAPA | 0.26 mM |
| oIMR8746 | 0.50 uM |
| oIMR8747 | 0.50 uM |
| oIMR9020 | 0.50 uM |
| oIMR9021 | 0.50 uM |
| Glycerol | 6.50 % |
| Dye | 1.00 X |
| Kapa 2G HS taq polym | 0.03 U/ul |
| DNA |
Cycling
| Step | Temp °C | Time | Note |
|---|---|---|---|
| 1 | 94.0 | -- | |
| 2 | 94.0 | -- | |
| 3 | 65.0 | -- | -0.5 C per cycle decrease |
| 4 | 68.0 | -- | |
| 5 | -- | repeat steps 2-4 for 10 cycles (Touchdown) | |
| 6 | 94.0 | -- | |
| 7 | 60.0 | -- | |
| 8 | 72.0 | -- | |
| 9 | -- | repeat steps 6-8 for 28 cycles | |
| 10 | 72.0 | -- | |
| 11 | 10.0 | -- | hold |
JAX uses a very high speed Taq (~1000 bp/sec), use cycling times recommended for your reagents.
JAX uses a 'touchdown' cycling protocol and therefore has not calculated the optimal annealing temperature for each set of primers.
Strains Using This Protocol
| Stock Number | Strain Name |
|---|---|
| 016198 | 129S6.Cg-Tg(Camk2a-tTA)1Mmay/JlwsJ |
| 003563 | B6.Cg-Tg(Cebpb-tTA)5Bjd/J |
| 003767 | B6.Cg-Tg(Eno2tTA)5021Nes/J |
| 003763 | B6.Cg-Tg(Eno2tTA)5030Nes/J |
| 005964 | B6.Cg-Tg(GFAP-tTA)110Pop/J |
| 002618 | B6.Cg-Tg(MMTVtTA)1Mam/J |
| 006232 | B6.Cg-Tg(Scgb1a1-rtTA)1Jaw/J |
| 034533 | B6.FVB-Tg(EmuSR-tTa)83Bop/DwfJ |
| 012433 | B6;C3-Tg(ACTA1-rtTA,tetO-cre)102Monk/J |
| 007678 | B6;SJL-Tg(KRT14-rtTA)208Jek/J |
| 006242 | C.Cg-Tg(Scgb1a1-rtTA)1Jaw/J |
| 006245 | C.Cg-Tg(SFTPC-rtTA)5Jaw/J |
| 013585 | FVB-Tg(Cdh5-tTA)D5Lbjn/J |
| 005625 | FVB-Tg(Pcp2-tTA)3Horr/J |
| 006222 | FVB.Cg-Tg(Scgb1a1-rtTA)1Jaw/J |
| 006225 | FVB.Cg-Tg(SFTPC-rtTA)5Jaw/J |
| 006209 | FVB.Cg-Tg(Tal1-tTA)19Dgt/J |
| 034802 | FVB/N-Tg(MMTV-tTA)25754Kuw/Mmjax |
| 004127 | FVB/N-Tg(Nes-rtTA)306Rvs/J |
| 005942 | FVB/N-Tg(Pf4-tTA/VP16)42Kra/J |
| 006875 | FVB/N-Tg(Tagln-rtTA)E1Jwst/J |
| 004937 | NOD.Cg-Tg(Ins2-tTA)1Doi/DoiJ |
| 005734 | NOD/Lt-Tg(Ins2-rtTA)1Ach/AchJ |
| 006999 | STOCK Dbttm1Geh Tg(Cebpb-tTA)5Bjd Tg(tetO-DBT)A1Geh/J |
| 024854 | STOCK Tg(Camk2a-tTA)1Mmay Fgf14Tg(tetO-MAPT*P301L)4510Kha/J |
| 003272 | STOCK Tg(Cebpb-tTA)5Bjd/J |
| 003273 | STOCK Tg(CMV-rtTA)4Bjd/J |
| 003271 | STOCK Tg(CMV-tTA)3Bjd/J |
| 005493 | STOCK Tg(Tek-rtTA,TRE-lacZ)1425Tpr/J |
| 008790 | STOCK Tg(tetO-DISC1*)1001Plet/J |
| 30 strains use this protocol | |