JAX Diversity Outbred Mice: A Genetically Diverse Mouse for a Diverse Human Population
Blog Post | August 11, 2020

The human population is incredibly diverse, with an infinite number of genomic variants that can determine both predisposition to disease and response to treatments. Therefore, it’s important to have a mouse model that encompasses the level of diversity seen in humans. The JAX Diversity Outbred (DO) mice have a random assortment of genetics from eight strains, resulting in a diverse population of genetically unique mice that more closely mimic the spectrum of diseases observed in the human population. Here is a brief look at this model.
Most traditional laboratory mouse strains were established by taking a mouse with unique characteristics and inbreeding it to produce a mouse population that is composed of virtually identical animals. This is what we call an inbred mouse strain. Inbred strains offer several advantages; they have well-defined genetics and characteristics, are reproducible, and can model specific human diseases. For example, the inbred strain A/J (JAX Stock #000646) is prone to several types of cancers, most notably adenocarcinoma, while the inbred NZO/HlLtJ (JAX Stock #002105) develops severe obesity (Figure 1). While a specific inbred strain may be valuable for a particular disease, this strain will not be suitable for all research applications and, therefore, must be chosen carefully
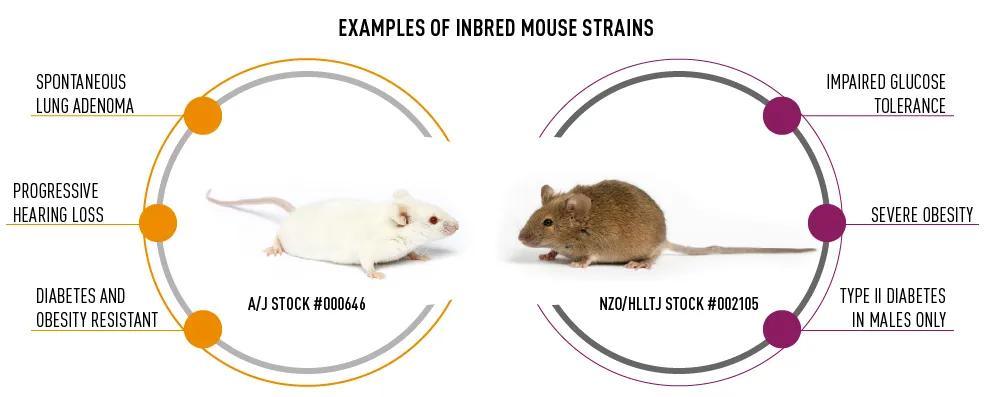
Figure 1 – Examples of inbred mouse strains and their unique phenotypes.
Testing a research hypothesis in a single inbred mouse strain is analogous of testing in a single human patient. Our human population is incredibly diverse, comprised of individuals with different genetics, ages, and sexes who are all in different environments. These intrinsic differences determine an individual’s predisposition to disease and their response to treatment. In a world where one size does not fit all, we need a diverse mouse model that can encompass the level of diversity in humans.
I’d like to introduce you to the JAX Diversity Outbred (DO) mouse population (009376). Diversity Outbred mice were generated by breeding eight founder inbred strains together to produce a population with a random assortment of genetics (Figure 2). The eight founder strains were chosen specifically because of their unique characteristics. For example, one develops type 1 diabetes (NOD/ShiLtJ), another is prone to cancer (A/J), and a third model develops obesity (NZO/HlLtJ). Together, these eight strains generate a diverse population of mice that can be used to model either susceptibility or resistance to a wide range of human diseases. Instead of choosing a specific inbred strain to test your hypothesis, you can do your research in a genetically diverse panel of mice that more closely mimics the spectrum of disease severity or resistance observed in the human population.
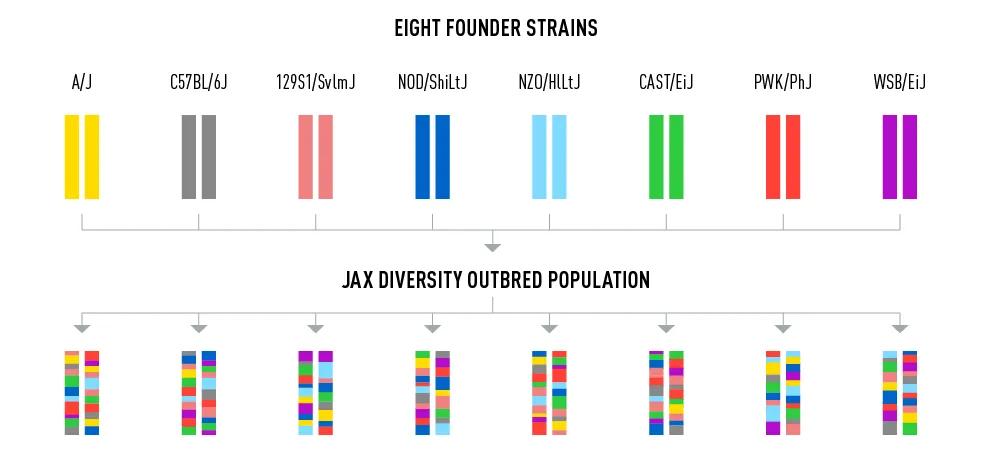
Figure 2 – Generation of the JAX Diversity Outbred (DO) mouse population. The inbred founder strains are shown as unique color-coded chromosomes. The eight strains were bred together in multiple steps to produce the Diversity Outbred population, which is composed of mice that are each a unique combination of genetics from the founders. (Figure is adapted with permission, from Collaborative Cross and Diversity Outbred Data Resources in the Mouse Phenome Database, Bogue et al. 2015).
So, what can you do with Diversity Outbred mice? Since the development of these mice in 2012, they have been used for research applications including pharmacology and toxicology, oncology, type II diabetes, infectious diseases, behavioral research, and more. In particular, the increased diversity in these mice promote:
- Investigation of the genetic and environmental basis of disease
- High-resolution mapping of genes and quantitative trait loci (QTL)
- Modeling drug and toxicant exposures in a diverse population
One early publication using Diversity Outbred mice highlights how these mice can simultaneously be a more accurate model for human toxicant exposures and determine genetic causes of susceptibility. Within this study, which was a collaboration between the National Institute of Environmental Health Sciences (NIEHS) and JAX, researchers exposed Diversity Outbred mice to several doses of benzene and measured benzene-induced chromosomal damage as an indicator of toxicity. They observed that Diversity Outbred mice were more susceptible to benzene-induced damage when compared to previous studies in a hybrid mouse strain. This is relevant to human health because benzene inhalation at lower doses than the established exposure limit has been shown to cause toxicity in humans. The exposure limits measured in Diversity Outbred mice more accurately modeled human response than the commonly used hybrid mouse model. Another promising finding of this study was that genetic mapping revealed a region in chromosome 10 that contained a pair of overexpressed sulfotransferases, which may be associated with toxicity from benzene.
The level of diversity seen in JAX Diversity Outbred mice unlocks many new research avenues. The real question is, how will you utilize Diversity Outbred mice to achieve your research goals? To learn more about JAX Diversity Outbred mice, visit the JAX DO datasheet and the recommended resources below.
References:
Bogue MA, Churchill GA, Elissa J Chesler. 2015. Collaborative Cross and Diversity Outbred data resources in the Mouse Phenome Database. Mamm Genome. 9-10:511-20. DOI: 10.1007/s00335-015-9595-6. [PMID: 26286858]
