Cre Lox Breeding for Beginners, Part 1
Blog Post | May 20, 2026
Simply put, the Cre/lox system rocks. It is one of the most powerful tools in the mouse geneticist’s toolbox. Why? Well, it enables them to generate tissue-specific and inducible knockouts and thereby have exquisite control over the location and timing of gene expression – important stuff when deleting a certain gene everywhere or during development leads to an embryonic lethal phenotype. And, it can be used to turn transgene expression on or off, track individual cells or cell lineages (as in the Brainbow or Confetti mice), generate inversions or translocations, and report gene expression. More novel and sophisticated uses for the Cre/lox system are being developed all the time. (Already know this? Here is Part 2!)
Research mouse models that incorporate the Cre/lox system typically involve generating double, triple, quadruple, or even n-tuple (is that really a word?) mutant/transgenic mice. As you can imagine, breeding such mice can get a wee bit complicated. Our Technical Information Services (TIS) is receiving more and more requests for help in designing breeding schemes for producing Cre/lox mice. Explaining these schemes can be difficult, and I swear that I can sometimes feel customers’ eyes cross while I’m trying to explain them over the telephone. So, in this post, I’m outlining a typical breeding scheme that I hope will help demystify the process of using the Cre/lox system to generate tissue-specific or inducible knockout mice. (In a future post, I will tackle the breeding schemes for using the system to turn transgenes on or off. I will also explain how to use Cre reporters).
Knockouts: remember, one cross just isn’t enough!
Here is the most efficient breeding scheme for generating Cre/lox tissue-specific or inducible knockouts. To generate mice that are heterozygous for a loxP-flanked allele and hemizygous/heterozygous for the cre transgene, mate a homozygous loxP-flanked mouse of interest to a cre transgenic mouse strain (see Figure 1 below). Approximately 50% of the offspring will be heterozygous for the loxP allele and hemizygous/heterozygous for the cre transgene.
Figure 1

Mate these mice back to the homozygous loxP-flanked mice (see Figure 2 below). Approximately 25% of the progeny from this mating will be homozygous for the loxP-flanked allele and hemizygous/heterozygous for the cre transgene. These will be your experimental mice.
Figure 2

About 25% of the offspring from this mating will be homozygous for the loxP flanked allele but will have no cre transgene. Depending on your experimental parameters, you can use these as controls (see Figure 3A below). Because the controls will not have Cre recombinase, any phenotypic differences between them and the Cre/lox mice should be due to the deleted gene in the Cre/lox mice. If appropriate for your experiments, you can maintain a colony that is homozygous for the loxP flanked allele, but in which one parent was hemizygous/heterozygous for cre and the other was a noncarrier/wild-type for cre.
This breeding scheme may not be the most efficient one for producing all Cre/lox strains. You will have to adapt it to the genotypes you want to produce and the genetic backgrounds and characteristics of the loxP-flanked and cre strains you use. To verify that Cre recombinase expression does not contribute to a phenotype of interest, I suggest using the cre strain itself (without any loxP-flanked alleles) as a control (Figure 3B). One of my colleagues, Andy, has written an awesome blog post on how Cre recombinase can produce a phenotype on its own. I highly recommend that you check out his blog.
Figure 3

The use of the Cre/loxP system for generating tissue specific or inducible knockouts is a powerful tool for mouse genetics, but it is only one piece of the total picture. Below, I would to focus on two additional issues: using Cre/lox mice to turn on or off transgenes, and using Cre reporter strains.
A Cre switch for transgenes
The Cre/lox system is one of the most powerful and versatile tools developed for mouse genetics. It gives mouse researchers sophisticated control over the location and timing of gene expression. Cre/lox is usually used to make knockout alleles, but it can also be used to activate gene expression.
The proper insertion of a loxP-flanked “stop” sequence (transcriptional termination element) between the promoter and transgene coding sequence blocks the expression of the gene. For a few examples of such strains, check out 008179 and 006481. Cre recombinase removes the stop sequence, so the transgene is expressed only in the cells where Cre is active (see Figure 4 below).
Figure 4
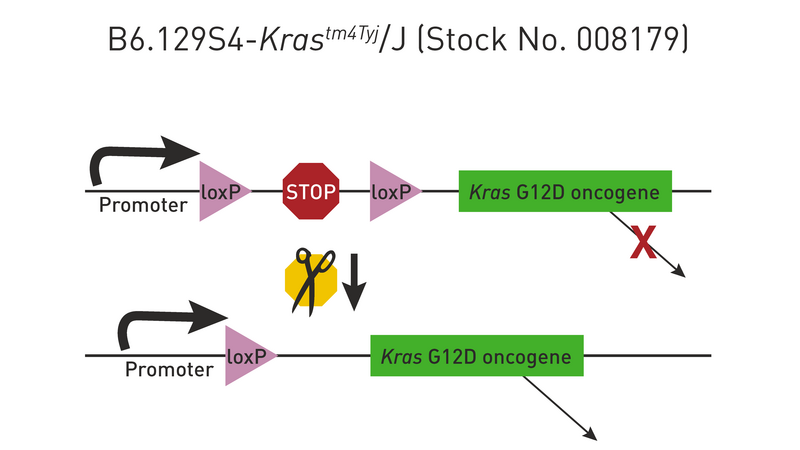
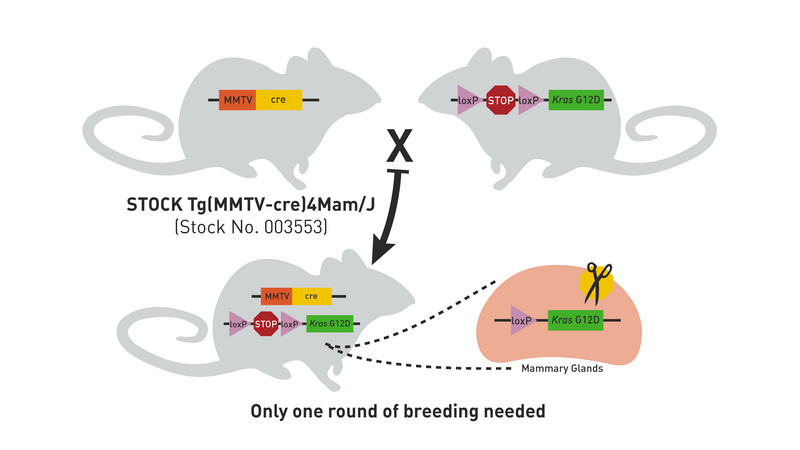
The Cre/lox system can also be used to produce strains in which a transgene is either inducible or expressed only in certain tissues. For example, mating the transgenic strain in Figure 1 to a strain that expresses Cre recombinase in mammary tissue produces double transgenic offspring that express the Kras oncogene only in the mammary glands (Figure 5). Notice that only one generation of breeding is required (Hooray!).
Figure 5
The Cre/lox system can be used to turn off a transgene. Such a transgene is designed with loxP sites flanking all or part of it. As an example, see 008917. In this example, expression of Cre recombinase removes the transgene (Figure 6).
Figure 6

If you are looking for the right Cre-expressing strain for your experiments, you can search for strains at the JAX Cre Repository or through the Cre Portal at MGI.
Extra! Extra! Read all about Cre expression!
When researchers started creating Cre recombinase mouse strains to generate tissue specific knockout mice, they needed an easy way to confirm that Cre recombinase was active only in certain tissues. This need led to the development of Cre reporter strains. These strains are designed to express a visible marker, such as green fluorescent protein (GFP) or LacZ, only after Cre recombinase excises a loxP-flanked stop sequence (Figure 7 and 8.)
Figure 7

When such a reporter strain is mated to a Cre-expressing strain, it produces offspring in which the visible marker is expressed only in tissues with Cre activity (Figure 8). Again, only one generation of breeding is required.
Figure 8

Cre reporter strains can also be used to generate a source of labeled tissues/organs, which allow researchers to track cells in transplant or adoptive transfer experiments.
Need help creating your Cre-lox model? JAX® Breeding Services can build experimental cohorts and colony controls from our expansive repository of Cre and loxP alleles, all maintained using high health standards.
