Let the experts in mouse genetics create a custom model to answer your scientific questions. With JAX, you can be confident your new model is precise and the genetic modification has been verified. We use a variety of technologies and strategies to deliver customized solutions that are specific to your needs.


JAX has worked with over 200 mouse strains for model generation projects. Base your model's genetic background on common strains like C57BL/6J or NSG™, or select from the 13,000+ strains in the JAX mouse collection.

Every project starts with a free consultation to assess the feasibility of your new model. Most custom projects will be supported by our no-risk guarantee: if we can't make your new mouse model, you will not pay.

Founder mice can be shipped directly to your institution or managed by JAX for colony expansion, preservation, characterization, testing, and analysis. All mice from JAX have a high health status and are specific-pathogen free (SPF).

Your dedicated project manager will provide regular updates, including founder and N1 data packets. Beyond model generation, turn your genetically modified JAX mouse into a validated model through expression analysis and/or phenotypic characterization.
Knock-out (KO) mice have a deletion or loss-of-function modification of your gene of interest. Both global and conditional knock-out models are typically made using CRISPR/Cas9 in mouse zygotes or ES cells. Researchers commonly use KO mice to:
A global knock-out deletes the mouse gene in all tissues throughout development and the mouse's lifetime. This approach is fast and efficient for studying gene function. JAX will delete one or more exons, resulting in a frameshift and a complete KO at the protein level in all tissues.
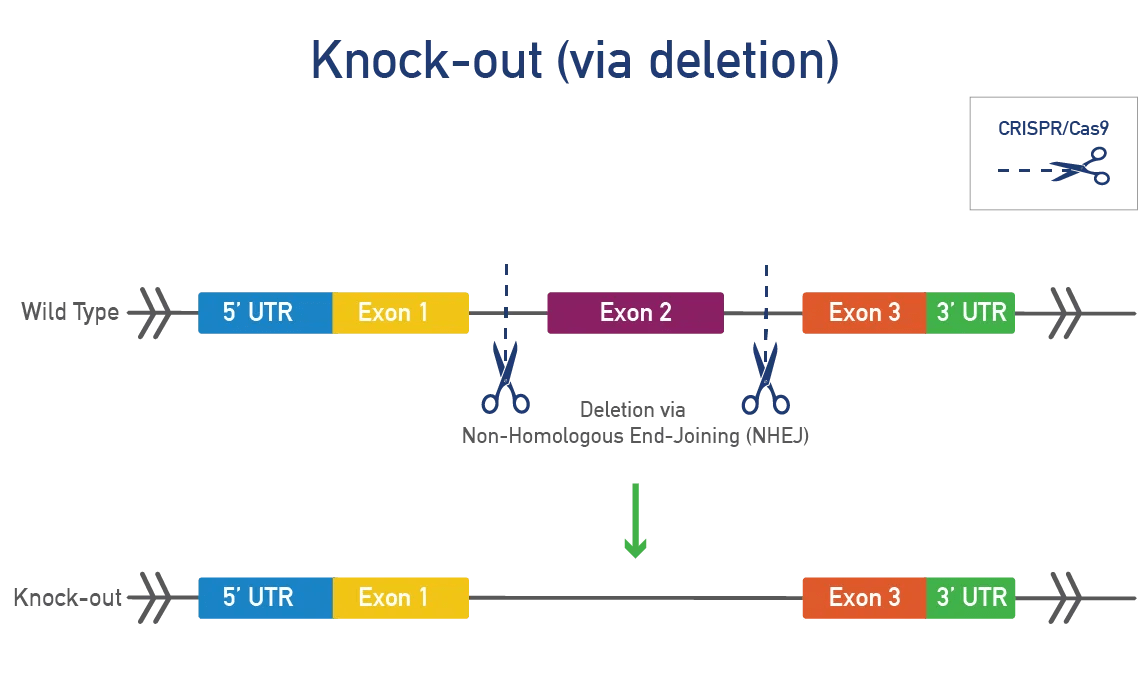
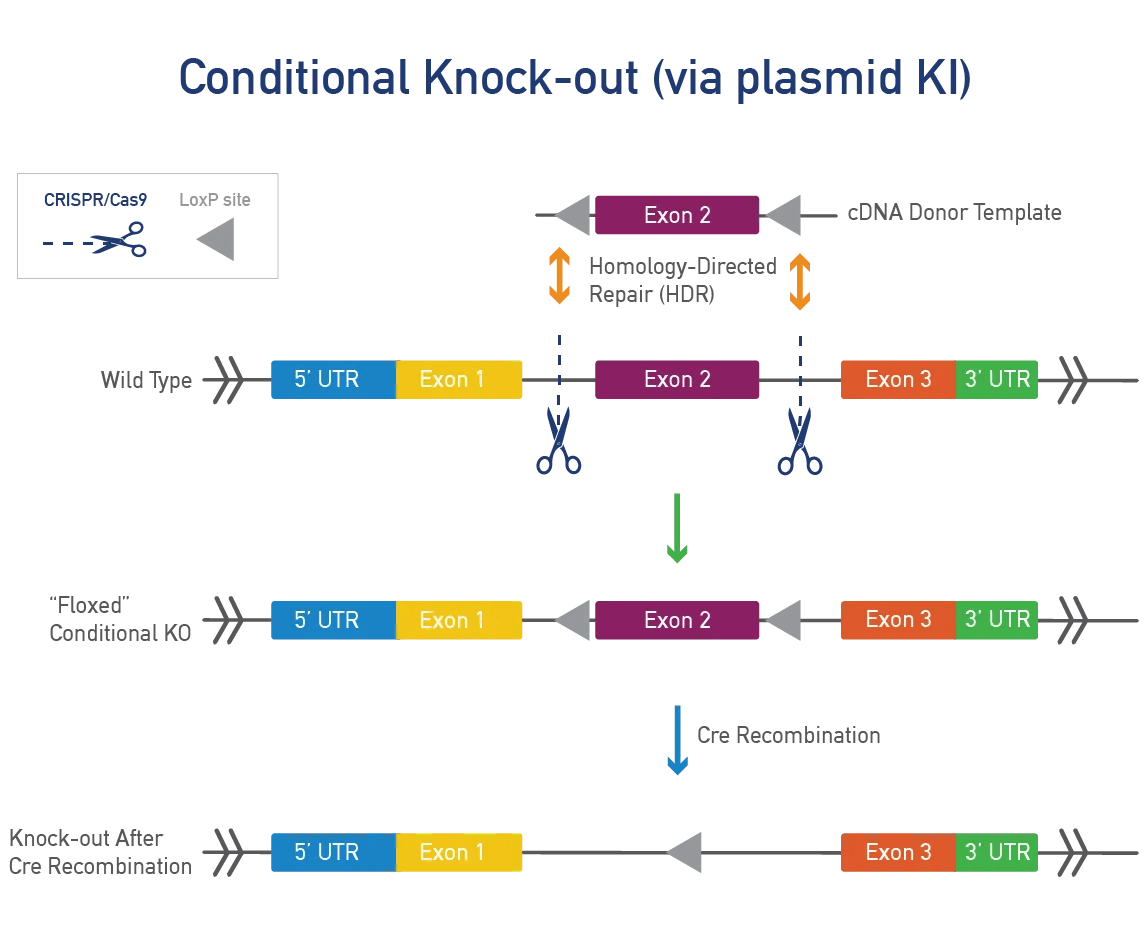
A conditional knock-out (cKO) can be bred to a recombinase-expressing strain to generate tissue-specific and/or inducible knock-outs. This approach is ideal for genes that produce severe phenotypes when disrupted. Most cKOs use the Cre-lox system, where LoxP sites are inserted on either side of a specific region of the mouse gene. When exposed to cre-recombinase, the region between the LoxP sites is removed, resulting in a functional KO. Alternative flanking sites or recombinases can also be inserted.
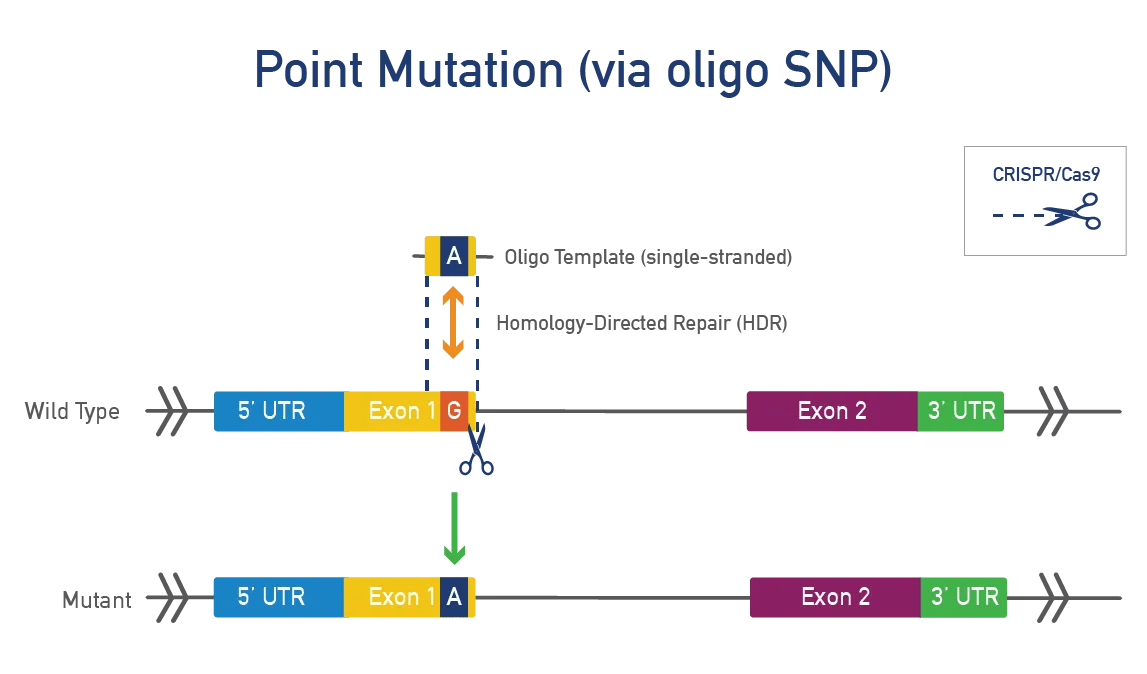
Point mutations occur when a single base pair is added, deleted, or changed. Researchers make point mutations in mice to model disease-causing mutations in humans. Point mutations are typically made using CRISPR/Cas9 in mouse zygotes or ES cells. These models allow researchers to:
A conditional point mutation allows expression of a mutation in specific tissues or time points, useful for severe phenotypes. A cre-dependent mutation needs cre-recombinase to be expressed. JAX uses a targeting vector with a LoxP-STOP-LoxP (LSL) cassette or dual LoxP system to control expression. We can also use Flp-FRT or Dre-Rox systems for generating conditional mutations.
A point mutation is made directly in the mouse gene of interest. This approach is precise, fast, and efficient and works best for genes that do not produce a severe phenotype when disrupted. JAX uses CRISPR/Cas9 to engineer these single-nucleotide changes. Single-stranded oligonucleotide (oligo) templates are used to make precise modifications while reducing the impact on surrounding genetic regions.
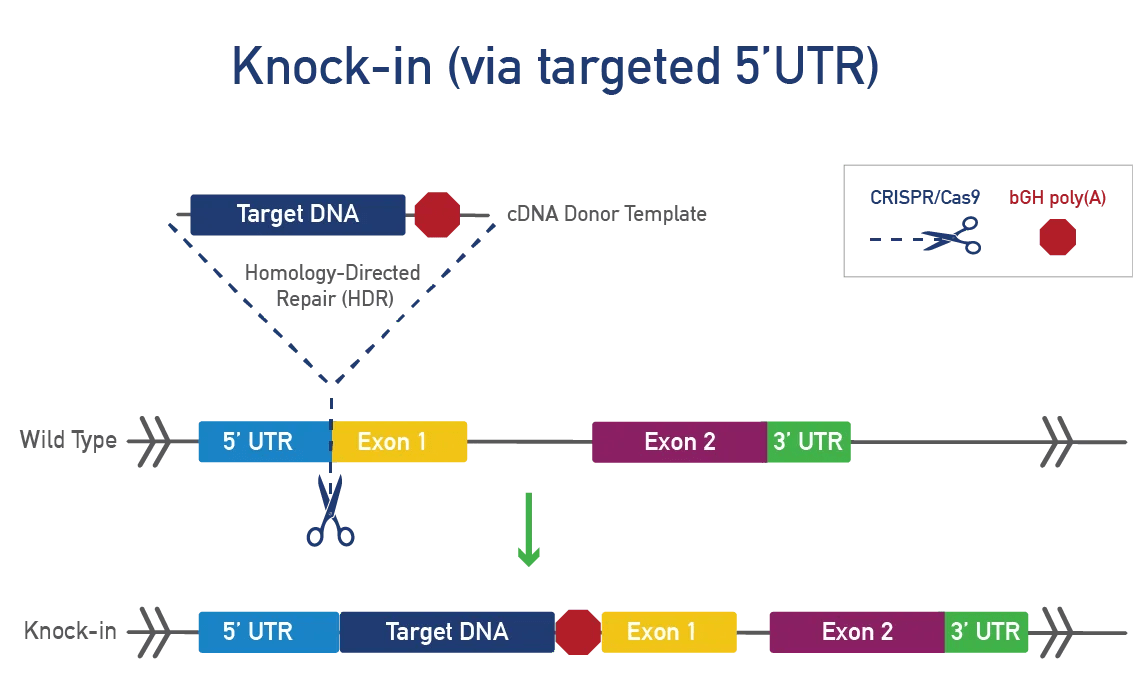
A knock-in involves inserting a DNA sequence into a specific mouse locus for various research purposes:
To generate a knock-in, CRISPR/Cas9 cuts DNA at a specific mouse locus and inserts a cDNA template using homology-directed repair (HDR). This method targets a gene or safe harbor locus, adds LoxP-STOP-LoxP or TetO for conditional expression, and incorporates tags or fluorescent proteins to track expression, ensuring precision and preventing unintended genetic consequences.
Translational modeling sometimes requires larger insertions. If your model exceeds CRISPR/Cas9 limits, JAX offers alternative techniques for larger insertions into the ROSA26 locus or traditional methods for random integration. These models allow researchers to study the role of a cluster of genes.
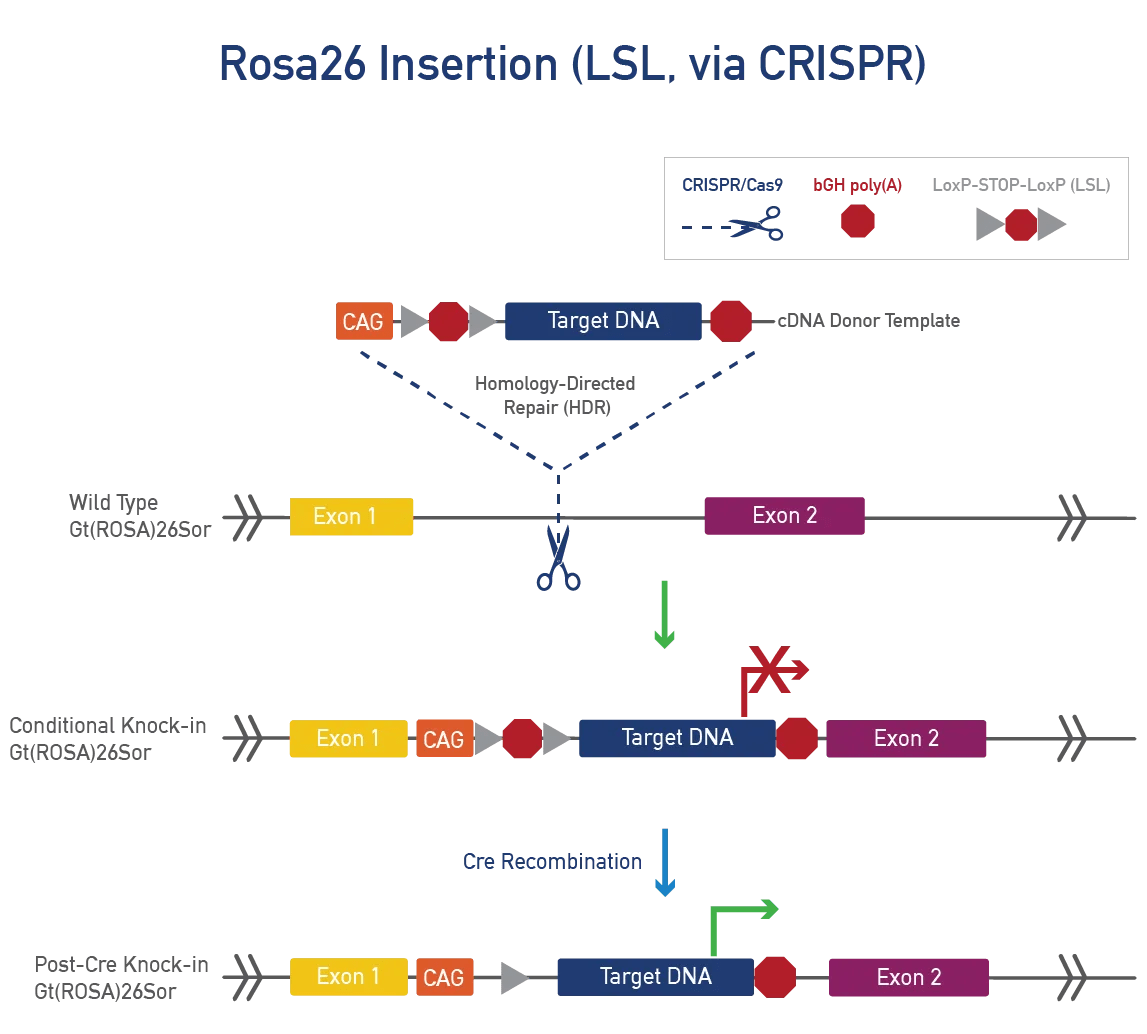
Gt(ROSA)26Sor (ROSA26) is a stable, safe-harbour locus with widespread expression. As a non-coding gene, it can be disrupted without phenotypic effects. ROSA26 is suitable for large, single-copy, targeted knock-ins using CRISPR/Cas9 or Bxb1 to insert cDNA donor templates. Promoters for widespread or tissue-specific expression can be added, as well as LSL cassettes for cre-dependent expression.
A transgene, such as a plasmid or BAC, is inserted randomly into the mouse genome. The size of random transgenics can range from less than 1 kb to several hundred kb with a BAC. This method suits large insertions when targeted insertion is not needed. To generate a random transgenic model, the DNA construct is synthesized or obtained and microinjected into pronuclear embryos. This produces multiple founder mice, each with potentially different integration sites, copy numbers, and expression levels.
CRISPR/Cas9 is a precise gene editing tool suitable for small genetic changes and larger modifications up to 4-6 kb. This method can be employed in many genetic backgrounds, eliminating the need for extensive backcrossing. JAX’s CRISPR/Cas9 Gene Editing Service includes designing and generating reagents, editing genes in mouse zygotes or ES cells, producing and sequencing founders, and one backcross to confirm heritability. Most projects have N1 animals ready within 6-9 months, depending on complexity. This service operates under licenses from The Broad Institute and Caribou Biosciences.
Bxb1 serine integrase is a new technology developed by JAX researchers that permits large, single-copy, targeted transgene insertions. If your construct exceeds the CRISPR size limits, you may consider using Bxb1 to insert your transgene into the ROSA26 safe harbor locus. Bxb1 has been reported to successfully make up to 40 kb insertions, and operationally JAX guarantees insertions up to 10 kb. This strategy is currently available on C57BL/6J, NOD/ShiLtJ, and NSG™ mouse models.
JAX can create plasmids or use your DNA constructs to generate transgenic mouse models, with a 90% success rate. The DNA Microinjection Service involves injecting transgenic DNA into pronuclear embryos, which are then transferred to pseudopregnant females. Founder mice are identified and shipped at six weeks old. Your model is typically ready in 13-15 weeks. Breeding Services can confirm germline transmission to the N1 generation.
JAX’s Embryonic Stem (ES) Cell Microinjection Service can generate new mouse models using your genetically modified mouse ES cell clones. ES cells are expertly injected into host blastocysts of your chosen genetic background. The resulting chimeric founder mice are identified by coat color at 6 weeks of age and ready to ship 13-15 weeks after receiving the ES cells. We also offer karyotyping of ES cells, microinjection of multiple ES cell clones, and breeding for germline transmission.