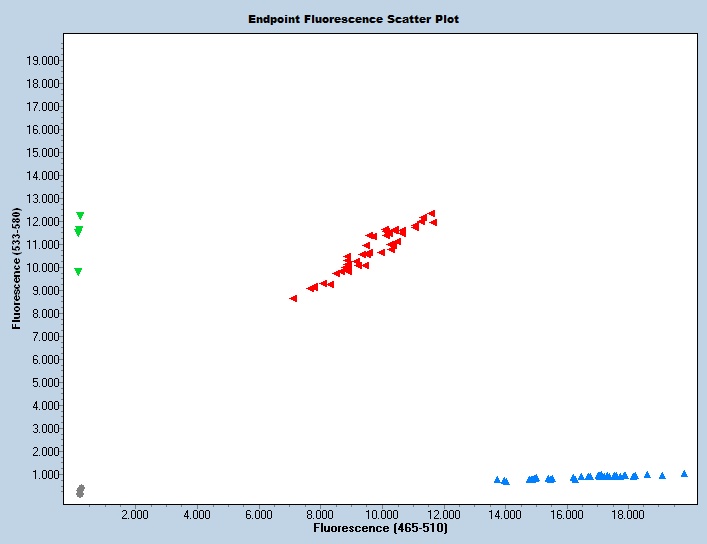
Notes
Taqman qPCR protocols are run on a real time PCR instrument. Use an appropriate instrument specific Fluorophore/Quencher combination. The transgene genotype is determined by comparing ΔCt values of each unknown sample against known homozygous and hemizygous controls, using appropriate endogenous references.
Expected Results
Mutant= 139 bp
Wild Type = 173 bp
Sequence
Wt Sequence: cccagagctgagcatcctaagcaagcactctgcgaactgagctacatcccagagttcataccaggatttaaggatctcaataggatagaatcaaaacagatactagtaagataaaaaccagtagtgatagaacggaagtcttgcttctaGAtaatagcatcttgccttcaaaaacttaactctgactatagagaacaaagacatcttagattcttaattcatgtg
Mutant Sequence:aaacagatactagtaagataaaaaccagtagtgatagaacggaagtcttgcttctagTGCGGCCGCACGTCTAAGAAACCATTATTATCATGACATTAACCTATAAAAATAGGCGTATCACGAGGCCCTTTCGTCTTCAAGAATTCCGATCATATTCAATAACCCTTAATataacttcgtataatgtatgctatacgaagttatTAGGTCCCTCGAAGAGGTTCACTAGataatagcatcttgccttcaaaaacttaactctgactatagagaaca
JAX Protocol
Protocol Primers
| Primer | 5' Label | Sequence 5' → 3' | 3' Label | Primer Type | Reaction | Note |
|---|---|---|---|---|---|---|
| 35320 | GTT AAG TTT TTG AAG GCA AGA TGC | Common | A | |||
| 35329 | Fluorophore-1 | CGG AAG TCT TGC TTC TAG ATA ATA GC | Quencher-1 | WT Probe | ||
| 36015 | CTG AGC ATC CTA AGC AAG CA | Wild type Forward | A | |||
| 36016 | CGA GGC CCT TTC GTC TTC | Mutant Forward | A | |||
| 36017 | Fluorophore-2 | AGG TCC CTC GAA GAG GTT CAC TAG ATA | Quencher-2 | MUT Probe |
Reaction A
| Component | Final Concentration |
|---|---|
| Kapa Probe Fast QPCR | 1.00 X |
| ddH2O | |
| 35320 | 0.40 uM |
| 36015 | 0.40 uM |
| 36016 | 0.40 uM |
| Wt Probe | 0.15 uM |
| Mutant Probe | 0.15 uM |
| DNA |
Cycling
| Step | Temp °C | Time | Note |
|---|---|---|---|
| 1 | 95.0 | -- | |
| 2 | 95.0 | -- | |
| 3 | 60.0 | -- | |
| 4 | -- | repeat steps 2-3 for 40 cycles | |
| 5 | 40.0 | -- | Forever |
Strains Using This Protocol
| Stock Number | Strain Name |
|---|---|
| 033964 | B6.129(Cg)-Ptf1atm2(cre/ESR1)Cvw Krastm4Tyj Ptentm1Hwu/J |
| 006440 | B6.129S4-Ptentm1Hwu/J |
| 013590 | B6.Cg-Tg(Tyr-cre/ERT2)13Bos Braftm1Mmcm Ptentm1Hwu/BosJ |
| 033760 | STOCK Braftm1Mmcm Gt(ROSA)26Sortm3(CAG-EYFP)Hze Nkx3-1tm4(cre/ERT2)Mms Ptentm1Hwu/AbshnJ |
| 033754 | STOCK Brca2tm1Brn Gt(ROSA)26Sortm3(CAG-EYFP)Hze Trp53tm1Brn Nkx3-1tm4(cre/ERT2)Mms Ptentm1Hwu/AbshnJ |
| 033752 | STOCK Gt(ROSA)26Sortm1(TMPRSS2/ERG)Key Nkx3-1tm4(cre/ERT2)Mms Ptentm1Hwu/AbshnJ |
| 033763 | STOCK Gt(ROSA)26Sortm2(myc*T58A)Rcse Nkx3-1tm4(cre/ERT2)Mms Ptentm1Hwu/AbshnJ |
| 033753 | STOCK Gt(ROSA)26Sortm3(CAG-EYFP)Hze Brca1tm2Cxd Nkx3-1tm4(cre/ERT2)Mms Ptentm1Hwu/AbshnJ |
| 033761 | STOCK Gt(ROSA)26Sortm3(CAG-EYFP)Hze Krastm4Tyj Nkx3-1tm4(cre/ERT2)Mms Ptentm1Hwu/AbshnJ |
| 033757 | STOCK Gt(ROSA)26Sortm3(CAG-EYFP)Hze Nkx3-1tm4(cre/ERT2)Mms Ptentm1Hwu Tg(ARR2/Pbsn-MYC)7Key/AbshnJ |
| 033751 | STOCK Gt(ROSA)26Sortm3(CAG-EYFP)Hze Nkx3-1tm4(cre/ERT2)Mms Ptentm1Hwu/AbshnJ |
| 033759 | STOCK Gt(ROSA)26Sortm3(CAG-EYFP)Hze Nkx3-1tm4(cre/ERT2)Mms Smad4tm2.1Cxd Ptentm1Hwu/AbshnJ |
| 033755 | STOCK Gt(ROSA)26Sortm3(CAG-EYFP)Hze Trp53tm1Brn Nkx3-1tm4(cre/ERT2)Mms Ptentm1Hwu/AbshnJ |
| 033756 | STOCK Gt(ROSA)26Sortm3(CAG-EYFP)Hze Trp53tm1Brn/Trp53tm3Tyj Nkx3-1tm4(cre/ERT2)Mms Ptentm1Hwu/AbshnJ |
| 14 strains use this protocol | |