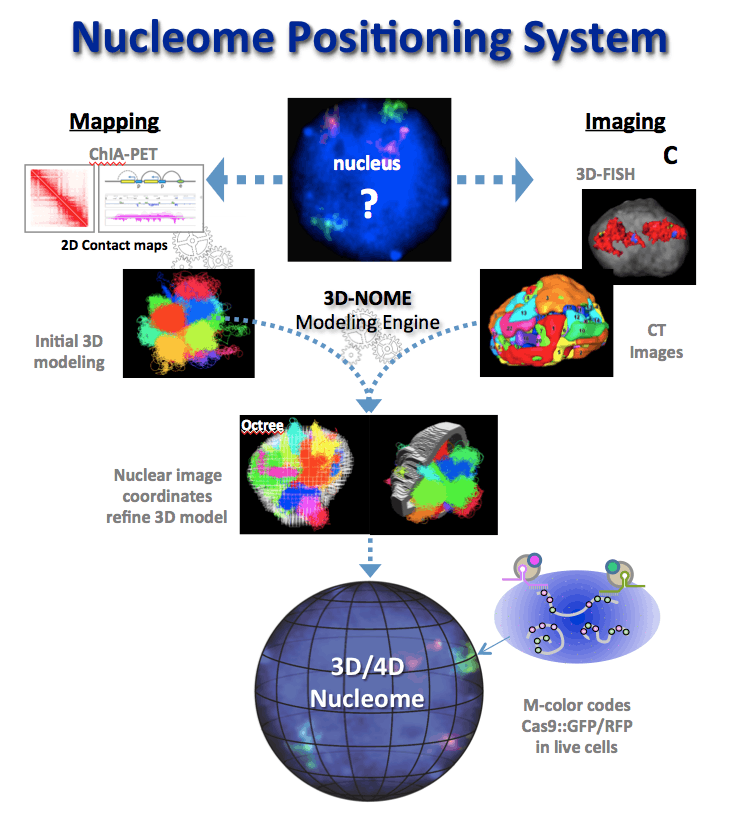
Although we typically think of the genome as a linear sequence, it is actually a dynamic three-dimensional structure. Gene loci and regulatory elements that are linearly distant—or even on separate chromosomes—may be brought in spatial proximity, and such interactions are of fundamental importance for understanding genome regulation.
The ultimate goal of the JAX 4D Nucleome Center is to deliver a Nucleome Positioning System (NPS) for the generation of complex maps of chromatin interaction network in the context of 3D genome structures. Such maps will be used to monitor and reference the dynamics of individual genomic elements, providing context to better understand gene function and the effects of genetic variation on gene function.
Our center is organized around four major goals:
- We will develop and improve technologies for high-resolution and haplotype-specific 3D genome mapping.
- We will refine an integrative and comprehensive data-analysis platform (3D-gNOME) for the study of nuclear architecture.
- We will elucidate the spatiotemporal regulation of 3D genome organization and function in human cells.
- We will use nuclear imaging and genome editing to structurally and functionally validate the 3D maps and predictions of regulatory interactions developed by the center.
Other Associated Researchers: Paul Blainey (Broad Institute)