For in-depth product & services help, ask our
Technical Information Scientists
Stock No: 007074
Protocol 29553: Standard PCR Assay - Cd40lg tm1 Imx (Tnfsf5) MCA
Version 2.2
Notes
The genotyping protocol(s) presented here have been optimized for reagents and conditions used by The Jackson Laboratory (JAX).
To genotype animals, JAX recommends researchers validate the assay independently upon receipt of animals into their facility.
Reaction cycling temperature and times may require additional optimization based on the specific genotyping reagents used.
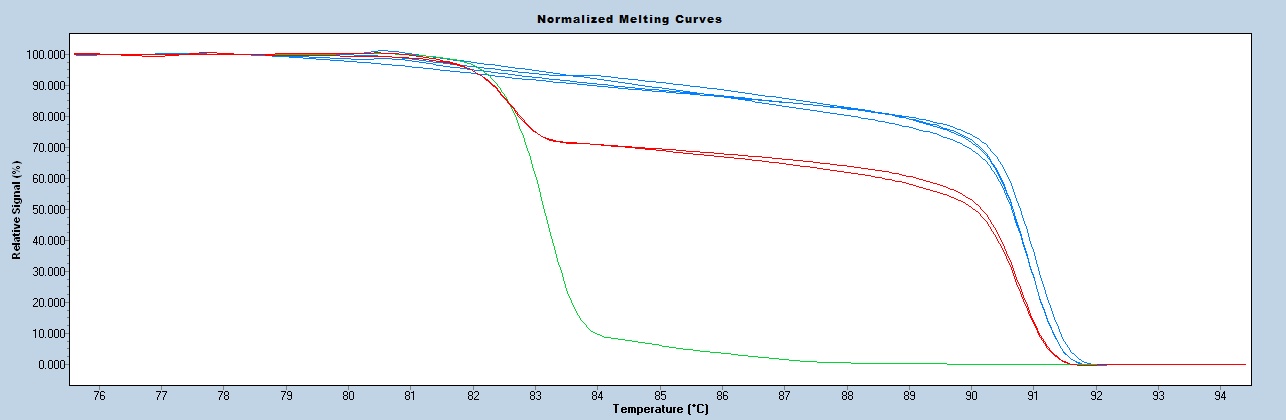
Expected Results
Mutant Tm = 89C
Wild type Tm = 80C
Wild type Tm = 80C
JAX Protocol
Protocol Primers
| Primer | 5' Label | Sequence 5' → 3' | 3' Label | Primer Type | Reaction | Note |
|---|---|---|---|---|---|---|
| oIMR0221 | GCC CTG AAT GAA CTG CAG GAC G | Mutant Forward | A | |||
| oIMR0222 | GGG TAG CCA ACG CTA TGT C | Mutant Reverse | A | |||
| oIMR0537 | GTT CCT CCA CCT AGT CAT TCA TC | Wild type | A | |||
| oIMR0538 | CCC AAG TGT ATG AGC ATG TGT GT | Wild type | A |
Reaction A
| Component | Final Concentration |
|---|---|
| ddH2O | |
| Kapa 2G HS buffer | 1.30 X |
| MgCl2 | 2.60 mM |
| dNTP KAPA | 0.26 mM |
| oIMR0221 | 0.50 uM |
| oIMR0222 | 0.50 uM |
| oIMR0537 | 0.50 uM |
| oIMR0538 | 0.50 uM |
| Glycerol | 6.50 % |
| Dye | 1.00 X |
| Kapa 2G HS taq polym | 0.03 U/ul |
| DNA |
Cycling
| Step | Temp °C | Time | Note |
|---|---|---|---|
| 1 | 94.0 | -- | |
| 2 | 94.0 | -- | |
| 3 | 65.0 | -- | -0.5 C per cycle decrease |
| 4 | 68.0 | -- | |
| 5 | -- | repeat steps 2-4 for 10 cycles (Touchdown) | |
| 6 | 94.0 | -- | |
| 7 | 60.0 | -- | |
| 8 | 72.0 | -- | |
| 9 | -- | repeat steps 6-8 for 28 cycles | |
| 10 | 72.0 | -- | |
| 11 | 10.0 | -- | hold |
JAX uses a very high speed Taq (~1000 bp/sec), use cycling times recommended for your reagents.
JAX uses a 'touchdown' cycling protocol and therefore has not calculated the optimal annealing temperature for each set of primers.
Strains Using This Protocol
This is the only strain that uses this protocol.